NanoDCAL
Who are the customers
using NanoDCAL?
Device engineers
Materials engineers
Academic researchers
Academic teachers
Benefits
NanoDCALPredict the electronic structure of virtually any material.
Accurately predict non-equilibrium properties of heterojunctions and devices.
Get the answer faster using NanoDCAL’s powerful implementation.
Using NanoDCAL is convenient and easy.
Download the NanoDCAL leaflet to get a summary of the software features!



Features
NanoDCALDescription
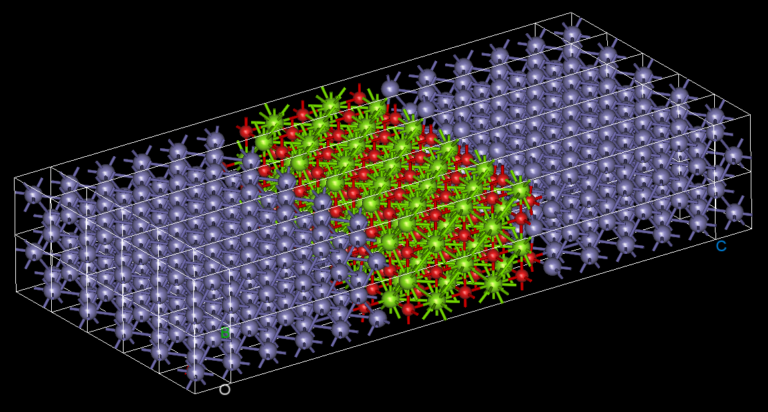
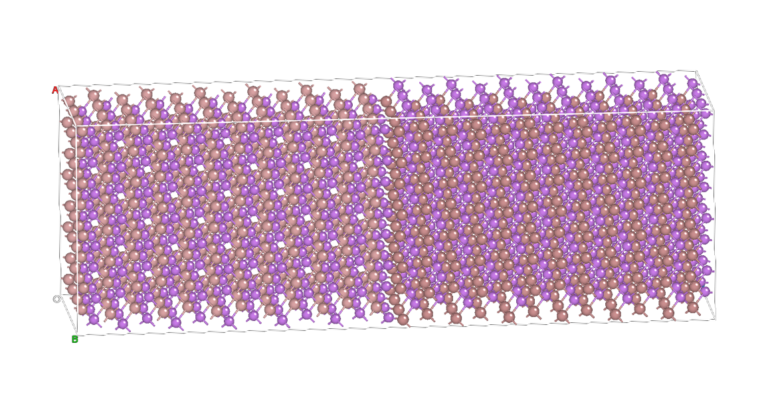
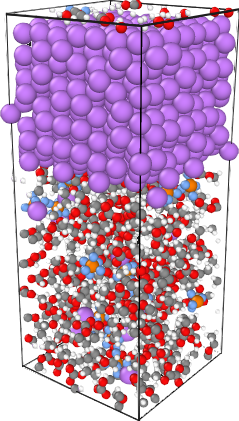
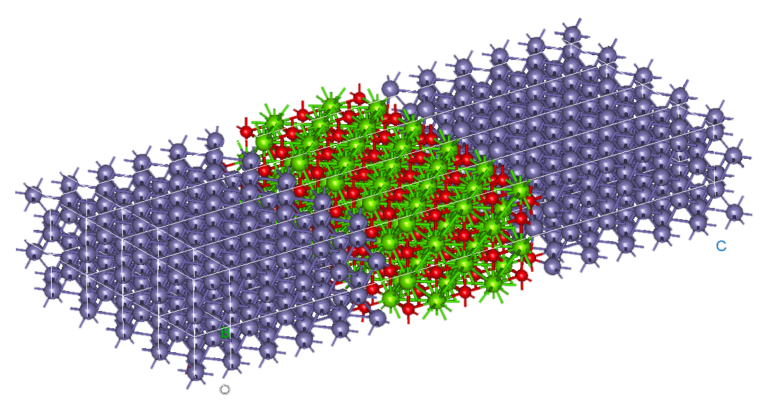
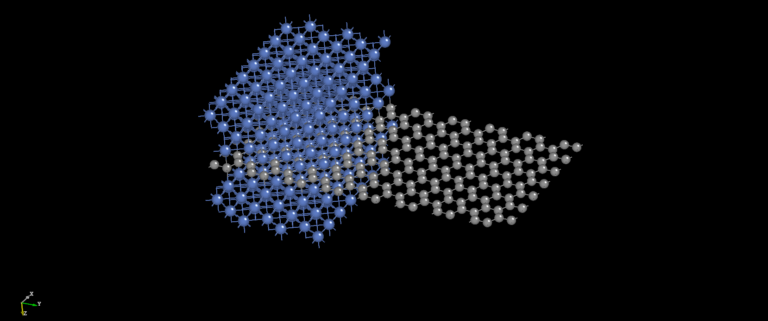
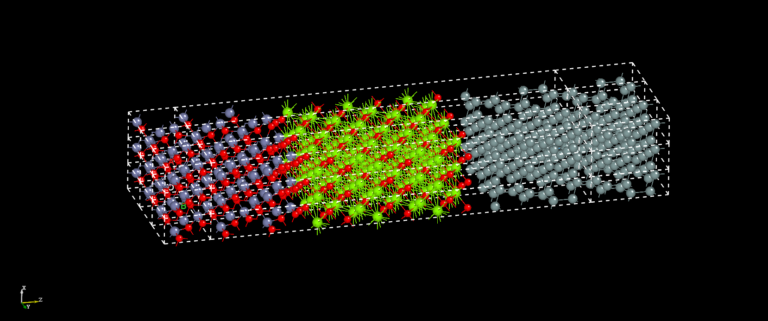
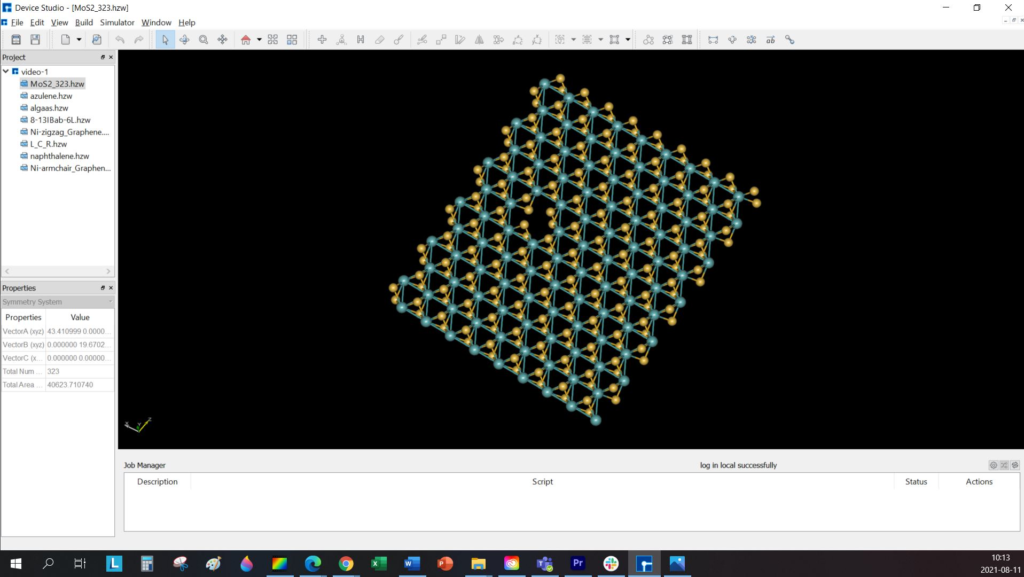
NanoDCAL offers reliable and powerful quantum transport simulation features to model nanostructures or nanodevices. It is an atomic orbital implementation of NEGF-DFT. It computes the Hamiltonian of materials and devices from first principles (i.e. without external parameters) using density functional theory (DFT) and simulates quantum transport phenomena within the Keldysh non-equilibrium Green function formalism (NEGF). NanoDCAL includes a large suite of methods for calculating important transport properties of your materials. NanoDCAL was used in hundreds of peer-reviewed papers in domains as varied as molecular electronics, nanotubes, topological insulators, batteries, magnetic tunnel junctions, metal grain boundaries (crystallites) and more. It has demonstrated efficiency, so why not test it? Unleash the full power and functionality of NanoDCAL by obtaining a parallel license.
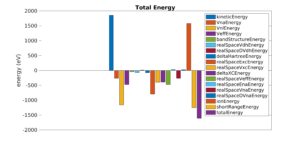
Ground state properties
Predict ground state properties like total energy, atomic forces, stress tensor.
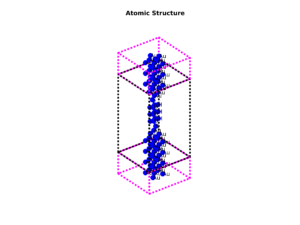
Atomic structure
Find equilibrium atomic positions and crystal shapes using NanoDCAL’s powerful structural optimizers.
Transmission & conductance
Get transmission channels and coefficients, conductance for bulk materials, surfaces and devices.
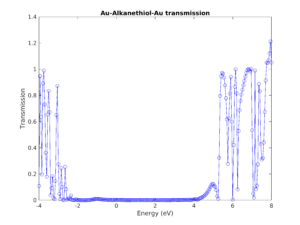
IV characteristic
Compute the current versus voltage characteristic to predict nanoscale device performance.
Multiple probes capability
NanoDCAL simulates devices involving more than two probes.
Atomic orbitals and pseudopotentials
Benefit from our accurate, efficient and complete database of atomic orbitals and pseudopotentials.
Fast & parallel solver
NanoDCAL is carefully optimized to get you the right answer faster. Key functions are written in C while most of the code is parallelized using both OpenMP and MPI.
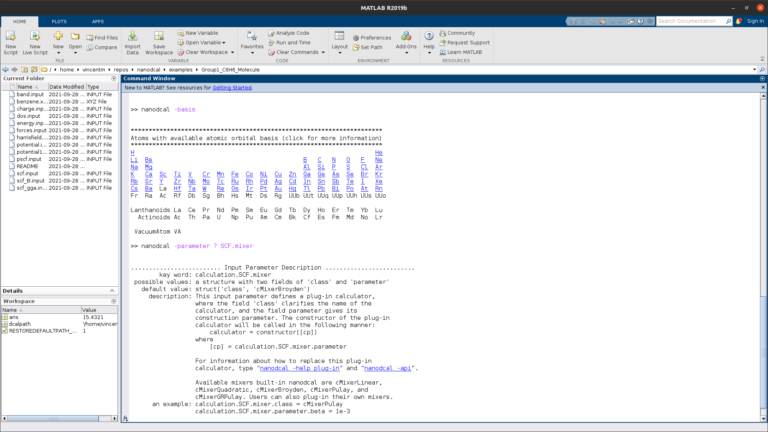
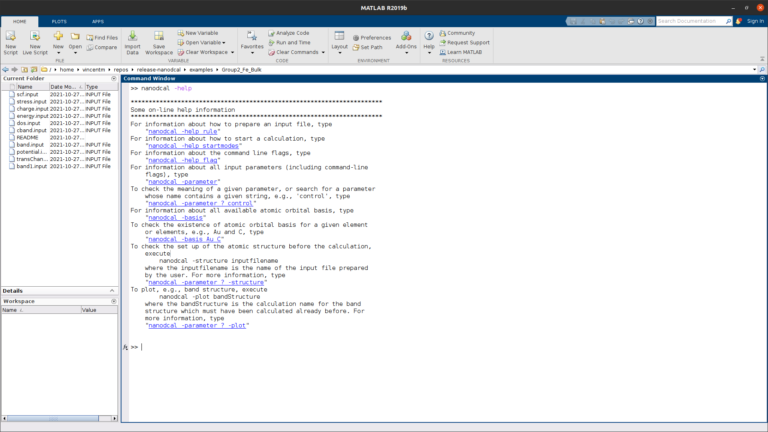
MATLAB integration
NanoDCAL is integrated in the MATLAB computing platform, allowing one to quickly and easily build workflows and visualize data.
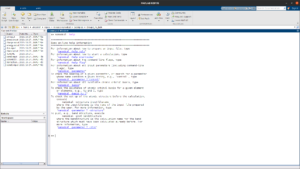
Spin & SOC
Include the physics of electronic spin and spin-orbit coupling via a state-of-the-art spin-DFT implementation (collinear and non-collinear formalisms).
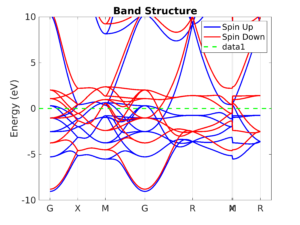
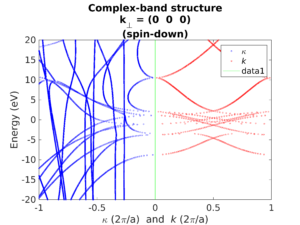
Spectral analysis
Understand the electronic structure of materials and devices with the band structure, complex band structure, (projected or total) density of states and more.
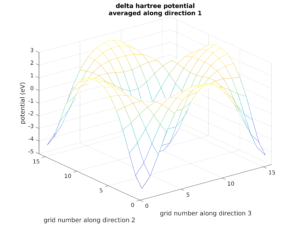
Real space analysis
Calculate and visualize relevant data in real space, such as local density of states, wavefunctions, scattering states and potentials.
Photocurrent
Compute the photocurrent of various materials and devices.
Thermal transport
Predict how thermal gradients drive electronic transport and vice versa.
What's new ?
A newly designed and enhanced version of NanoDCALhas just been released: NanoDCAL+. Written in Python and Fortran, it provides new Python calculators and features, more performance and scalability for workstations and supercomputers alike. It is a complementary version of NanoDCAL. Please see the NanoDCAL+ product page for more information!
Current NanoDCAL version is 2022B (v3.1.0).
NanoDCAL user documentation
To access to NanoDCAL user documentation, click the link below: installation and user manuals, tutorials, theory and technical information pages are all there for your support and to get you up to speed as quickly as possible.